FlyBrainLab
FlyBrainLab is an open-source interactive computing platform for studying the function of executable circuits constructed from fruit fly brain morphology data. The workflow of the FlyBrainLab platform is designed with three main capabilities in mind that are schematically depicted in the figure below:
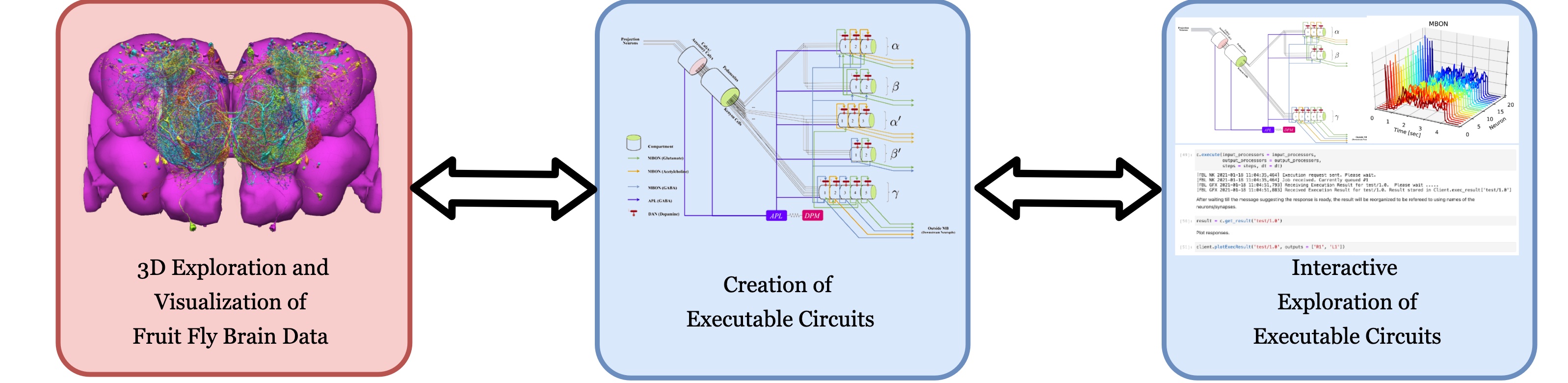
- (left) 3D exploration and visualization of fruit fly brain morphology data,
- (middle) creation of executable circuits directly from the explored and visualized fly brain morphology data, and
- (right) interactive exploration of the functional logic of the devised executable circuits.
In the video below the workflow supported by FlyBrainLab for analyzing, evaluating and comparing circuit models of the fruit fly Central Complex (CX) is demonstrated with the FlyCircuit dataset. Three of the circuit models published in the literature were implemented. The interactive capabilities of the three models are shown side-by-side, including the visualization of the morphology of CX neurons and the corresponding executable circuits. Finally, an executable neural circuit for each model was instantiated and executed, and the results displayed in parallel.
More of the details of the methodology of comparing the computational models of the CX devised by three different research groups using FlyBrainLab can be found in
1. Aurel A. Lazar, Tingkai Liu, Mehmet K. Turkcan, and Yiyin Zhou, Accelerating with FlyBrainLab the Discovery of the Functional Logic of the Drosophila Brain in the Connectomic and Synaptomic Era, eLife , February 2021.
The Architecture of the FlyBrainLab
The main user-side and server-side components of the FlyBrainLab are depicted in the figure below.
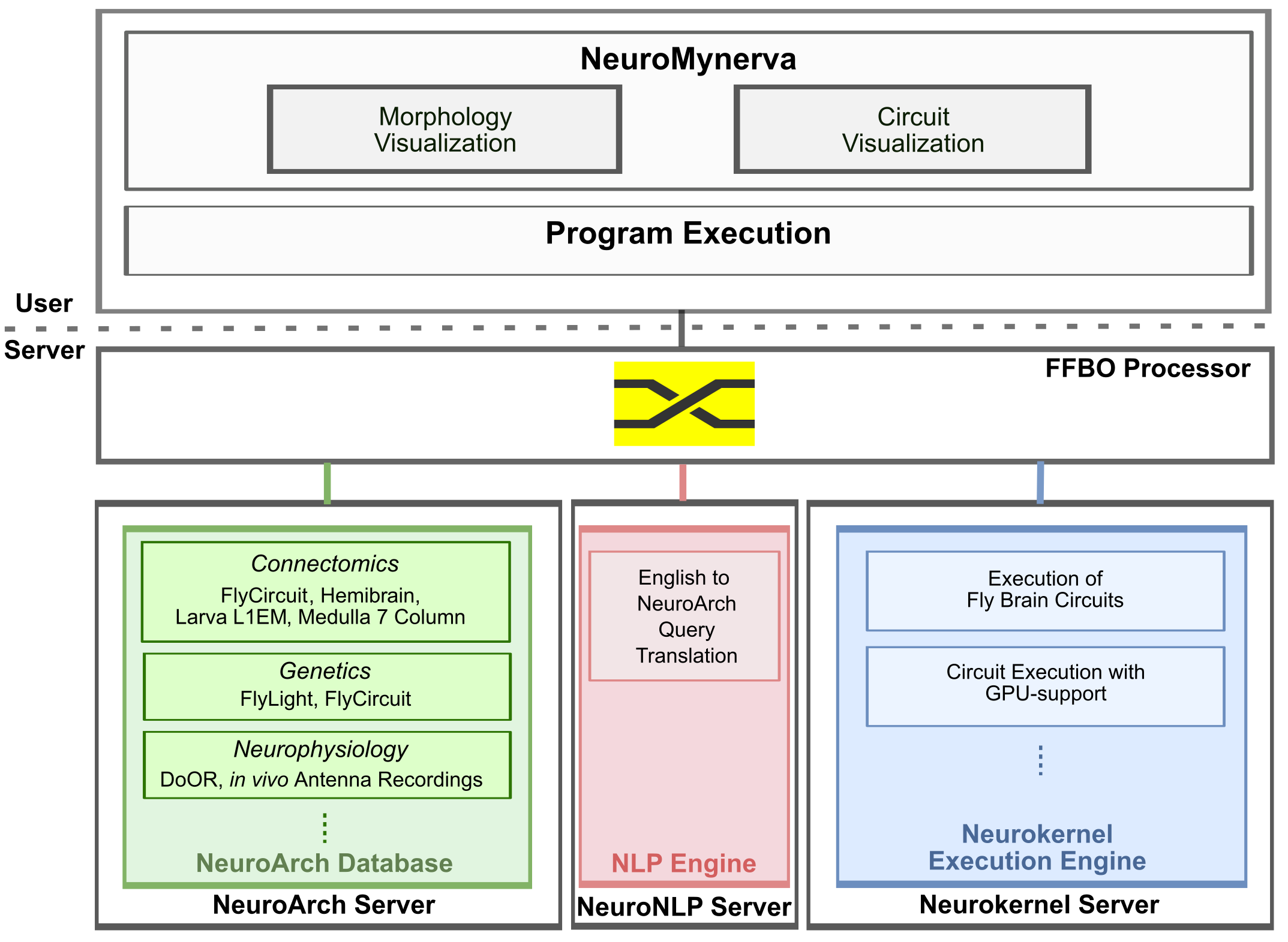
The FlyBrainLab user-side components consist of the NeuroMynerva User Interface and the FlyBrainLab Client back-end (Program Execution). The former provides capabilities such as morphology visualization and interactive circuit diagrams, and the latter provides program execution for communicating with the server components. FlyBrainLab Code Repository.
A description of the main components of the architecture of FlyBrainLab can be found in
1. Aurel A. Lazar, Tingkai Liu, Mehmet K. Turkcan, and Yiyin Zhou, Accelerating with FlyBrainLab the Discovery of the Functional Logic of the Drosophila Brain in the Connectomic and Synaptomic Era, eLife , February 2021.
2. Aurel A. Lazar, Mehmet Kerem Turkcan, and Yiyin Zhou, A Programmable Ontology Encompassing the Functional Logic of the Drosophila Brain, Frontiers in Neuroinformatics, April 2022.
NeuroMynerva User Interface
NeuroMynerva is the user interface of FlyBrainLab. It is a browser-based application that substantially extends upon JupyterLab to combine the functionality of NeuroNLP for 3D visualization and exploration of fly brain data, and of NeuroGFX for interactively exploring the function of executable neural circuits using interactive circuit diagrams. Both can interoperate with code execution in a Jupyter notebook. NeuroMynerva Code Repository.
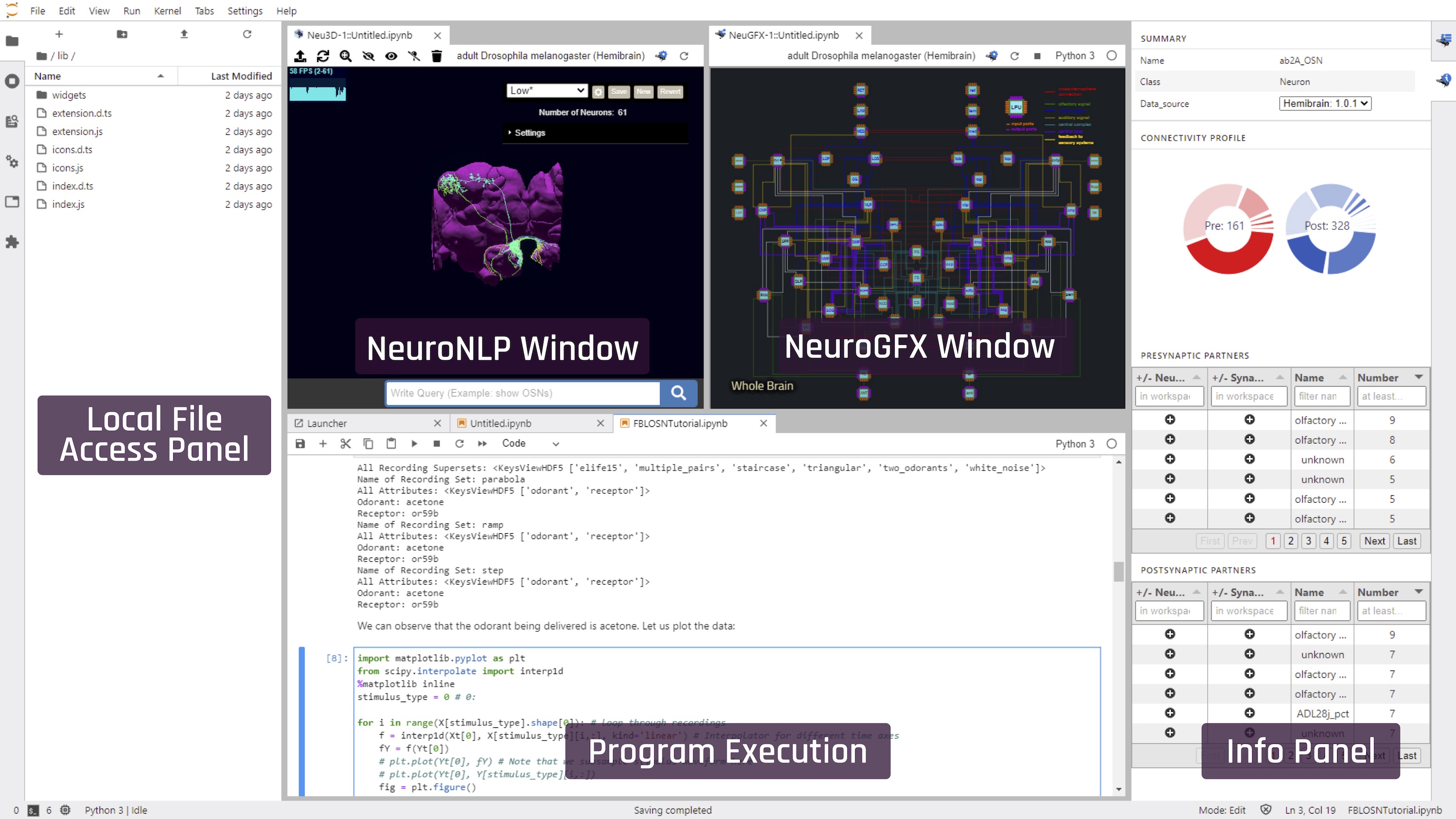
For a description of the UI see
1. Aurel A. Lazar, Tingkai Liu, Mehmet K. Turkcan, and Yiyin Zhou, Accelerating with FlyBrainLab the Discovery of the Functional Logic of the Drosophila Brain in the Connectomic and Synaptomic Era, eLife , February 2021.
NeuroArch Database
NeuroArch is the first graph database that provides a novel data model for representation and storage of connectomic, synaptomic, cell type, activity and genetic data of the fruit fly brain with cross-referenced executable circuits. The NeuroArch data model is the foundation of the integration of fruit fly brain data and executable circuits in FlyBrainLab. NeuroArch Code Repository.
- Lev E. Givon, Aurel A. Lazar and Nikul H. Ukani, Neuroarch: A Graph-Based Platform for Constructing and Querying Models of the Fruit Fly Brain Architecture, Frontiers in Neuroinformatics , Number 42, August 2014, Leiden, The Netherlands.
- Lev E. Givon, Aurel A. Lazar and Nikul H. Ukani, NeuroArch: A Graph dB for Querying and Executing Fruit Fly Brain Circuits, Neurokernel Request for Comments, Neurokernel RFC #5, December 2015.
Neurokernel Execution Engine
Neurokernel is the first GPU-based execution engine for emulating the fruit fly brain. It provides a comprehensive API ensuring that independently developed brain circuit models of different brain regions are interoperable and can be combined into a whole brain emulation. Neurokernel Code Repository.
- Lev E. Givon and Aurel A. Lazar, Neurokernel: An Open Source Platform for Emulating the Fruit Fly Brain, Neurokernel Request for Comments, Neurokernel RFC #4, October 2015.
- Lev E. Givon and Aurel A. Lazar, Neurokernel: An Open Source Platform for Emulating the Fruit Fly Brain, PLOS ONE 11(1): e0146581, doi:10.1371/journal.pone.0146581, January 2016.
The Bionet Group is supported by grants from
 |
 |
 |
 |